The SARS-CoV-2 genome contains 28 genes that encode 16 non-structural proteins (Nsp), 4 structural proteins (N, E, S, M) and 8 accessory proteins (Orf, open reading frame).
Orf6, Nsp6 and Orf7a induced the highest toxicity when over-expressed in human HEK 293 T cells. All three proteins showed membrane localization in COS-7 cells (fibroblast-like cell lines derived from monkey kidney tissue). Orf6 is most cytotoxic and localized to the endoplasmic reticulum, autophagosome (deliverinh cytoplasmic components to the lysosomes) and lysosomal membranes. Lee et all identified seven SARS-CoV-2 proteins capable of inducing significant cell viability defects. Among these seven, SARS-CoV-2 Orf6 displayed the strongest toxicity phenotype with near 50% loss in cell viability, followed by the Nsp6 and Orf7a proteins which each showed ~ 30–40% reduction in cell viability. Four other proteins (Nsp13, Nsp14, Orf3a and M) induced milder yet significant cell death, whereas the other 21 proteins did not induce detectable cellular toxicity.
In synthesizing a protein, gene (in the nucleus) must be activated and the instructions encoded in the gene’s DNA must be converted to RNA. This mRNA then be transported out of the nucleus to ribosomes, then it “read” the mRNA and begin assembling the protein.
Two proteins, an mRNA export factor called Ribonucleic Acid Export 1 (Rae1), and a nuclear pore complex protein, called (Nup98), play a key role by binding together with the newly synthesized mRNA and shepherding it through the nuclear membrane pore.
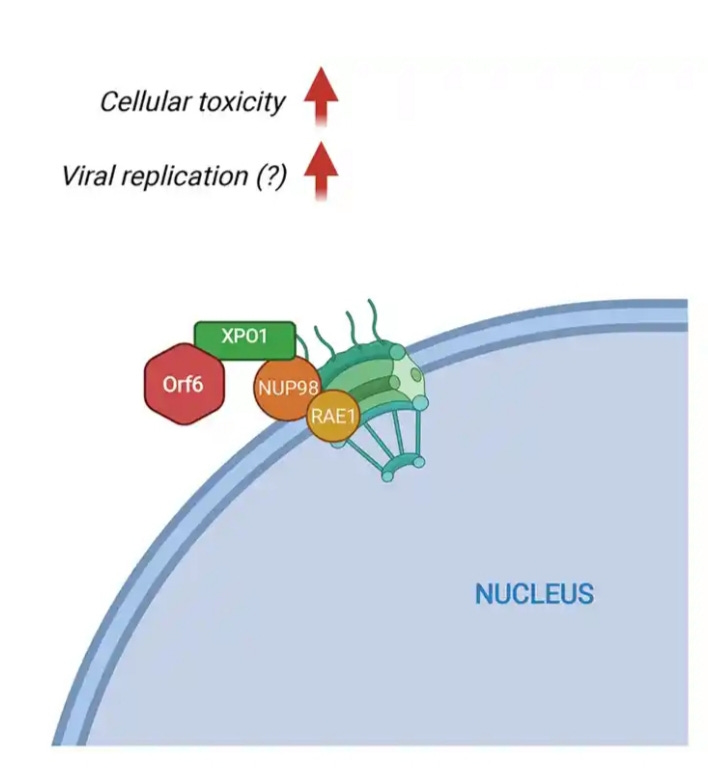
Orf6 inhibits infected cells’ ability to produce proteins needed to respond to infection. When cells were infected with SARS-CoV-2, Orf6 bound to and disabled the Rae1-Nup98 complex. Orf6 localizes at the nuclear pore complex (NPC) and directly interacts with Nup98-Rae1 via its C-terminal domain to impair docking of cargo-receptor (karyopherin/importin) complex and disrupt nuclear import.
Orf6 prevents a broad range signaling proteins from entering the nucleus (by clogging it), including signaling proteins that make the cell incapable to send interferons (IFNs). IFNs are a family of glycoproteins with essential roles in restricting viral replication and in modulating the antiviral immune response.
Orf6 blocks the nucleocytoplasmic transport. This results in giving the virus time to replicate before the host shows any symptomps. Thus, the person can transmit the virus though that person doesn't show any symptomps. Orf6 has a big role in creating asymptomatics Covid-19.
B.1.525 is the only variant of SARS-COV-2 known with Orf6 nonsynonimous mutation. Nonsynonymous mutations have a much greater effect on an individual than a synonymous mutation. This variant was reported in :
US (December 2020) 558 sequences.
UK (January 2021) 338 sequences.
Germany (January 2021) 347 sequences.
France (January 2021) 275 sequences.
Italy (February 2021) 162 sequences.
Denmark (January 2021) 121 sequences.
Norway (January 2021) 69 sequences.
The rest 40 countries have less than 50 sequences, varying from 1-49 sequences.
B.1.525 carries mutations found in UK, South Africa, and Brazil variants. Based by E484 mutation, beside reducing convalescent serum neutralization and monoclonal antibodies, it also causes a small but significant reduction in neutralization by mRNA Covid vaccines.
B.1.525 is a variant that more likely can create a bigger asymptomatics numbers in Covid-19 transmission. A greater effort in sequencing SARS-CoV-2 genome should be considered for every country. Though it’s not easy, analyzing the representative asymptomatic data in a region will predict the transmission outcome and the positive rate. Thus, any prophylactic drugs and supplements (e.g. Vitamin C & D, Zinc) should be considered served on daily basis.
Sources:
https://newsroom.uw.edu/news/how-one-sars-cov-2-protein-keeps-cells-fighting-back
https://www.pnas.org/content/117/45/28344
https://covariants.org/variants/20A.S.484K
https://cellandbioscience.biomedcentral.com/articles/10.1186/s13578-021-00568-7
https://www.thoughtco.com/synonymous-vs-nonsynonymous-mutations-1224600
Picture:
https://cellandbioscience.biomedcentral.com/articles/10.1186/s13578-021-00568-7